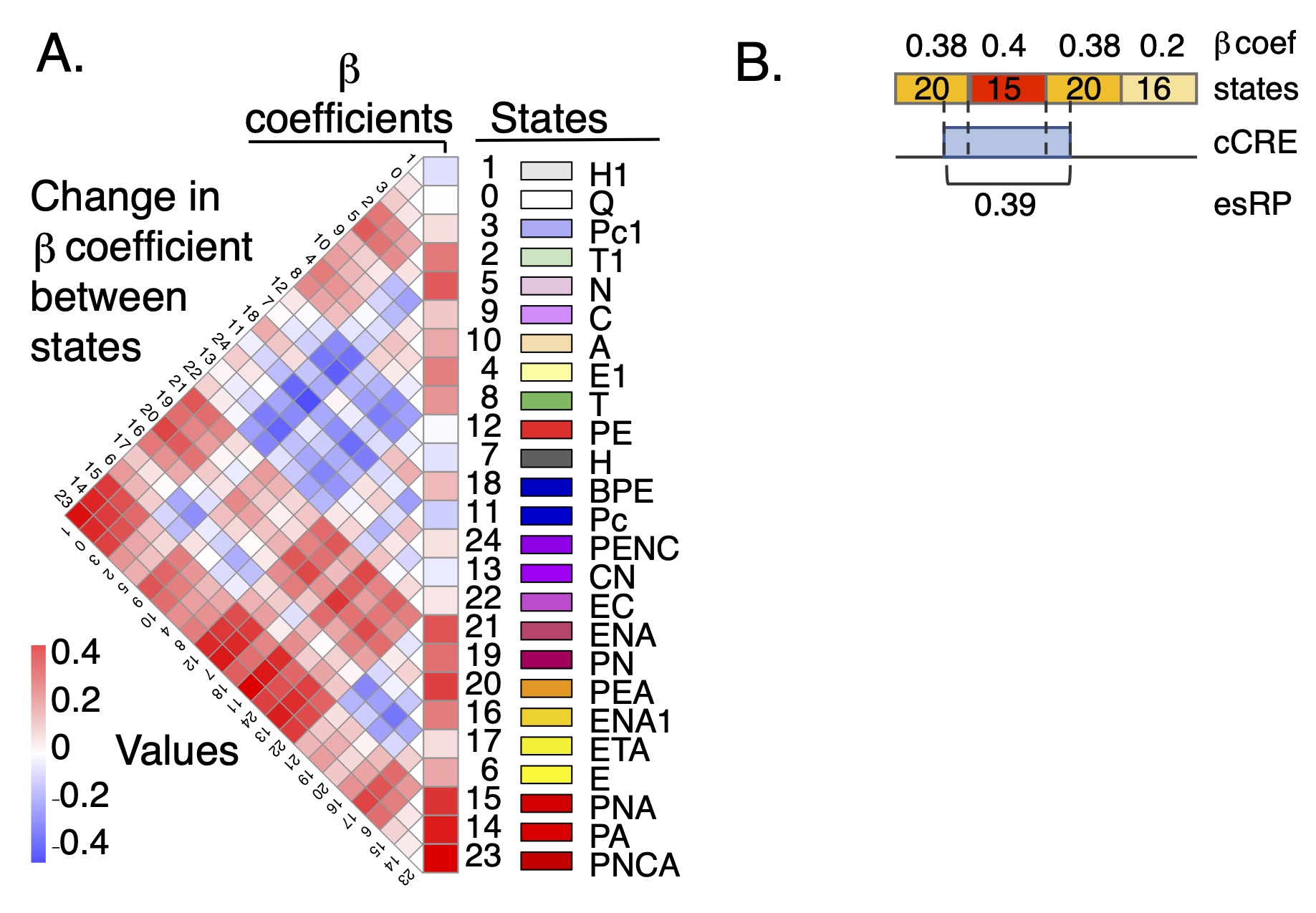
esRP scores for cCREs The epigenetic state Regulatory Potential (esRP) score is an estimate of the contribution of a candidate cis-regulatory element, or cCRE, to the level of expression of a (potential) target gene. The score is derived from the beta coefficients that relate the proportion of cCRE intervals in a particular multi-feature epigenetic state (available in the super track Epigenetic states) to expression levels of target genes in a multivariate linear regression (Xiang et al. 2024; Figure 1). The esRP score is simply a weighted sum of beta coefficients of states that cover a cCRE in a cell type, where the weights are the region covered by different states. The cCREs were defined as DNA intervals with a high signal for chromatin accessibility in any cell type (Xiang et al. 2020 and 2024). They can be visualized using the companion super track Candidate CREs, and they are described more thoroughly on the Track Settings page of that super track.

The most useful setting is "pack" mode, in which the cCREs are labeled with the value of the esRP scores in each cell type. In "dense" mode, the display simply shows the position of each cCRE.
The track names (short name and the end of the long name) give an abbreviation for the blood cell type and replicate number (r1 or r2).
Mouse primary blood cells purified predominantly using cell surface markers include: LSK = Lin-Sca1+Kit+ cells from mouse bone marrow containing hematopoietic stem and progenitor cells, CMP = common myeloid progenitor cell, MEP = megakaryocyte-erythrocyte progenitor cell, ERY = erythroblast, GMP = granulocyte monocyte progenitor cell, MON = monocyte, NEU = neutrophil, CLP = common lymphoid progenitor cell, B = B cell, NK = natural killer cell, T_CD4 = CD4+ T cell, T_CD8 = CD8+ T cell, CFUE = colony forming unit erythroid, fl = designates ERY derived from fetal liver, ad = designates ERY derived from adult bone marrow, CFUMK = colony forming unit megakaryocyte, iMK = immature megakaryocyte, MK_fl = megakaryocyte derived from fetal liver. AVE is a track with state assignments based on the average signal for each epigenetic feature across cell types.
Data from several immortalized cell lines were included. The G1E cells are an immortalized, GATA1-null cell line derived from mouse embryonic stem cells by gene targeting; these cells proliferate in culture as immature erythroid progenitor cells (Weiss, Yu, Orkin 1997). A stable subline of these cells, called G1E-ER4, undergoes terminal erythroid maturation when GATA1 function is restored as an activatable fusion of GATA1 to the ligand-binding domain of the estrogen receptor (ER). Untreated G1E-ER4 cells, carrying the inactive GATA1-ER, proliferate without differentiation, but treatment with estradiol (E2) activates the hybrid protein, effectively complementing the GATA1 loss-of-function and allowing synchronous erythroid differentiation and maturation (Gregory et al. 1999). An additional cell line model used here are murine erythroleukemia (MEL) cells, which can be chemically induced to mature into erythroblast-like cells with increased hemoglobin (iMEL). HPC7 cells are an immortalized line that serves as a model for mouse hematopoietic progenitor cells (Pinto do O 2002). These cells are capable of differentiation in vitro into more mature myeloid cells. CH12 cells are an immortalized line that is a model for mouse B cells; the epigenetic data on CH12 cells were used to generate the B cell epigenetic state annotation.
The methods for multivariate linear regression of epigenetic states versus expression levels to estimate beta coefficients and the calculation of esRP scores are described on the Track Settings page of the parent super track Epigenetic state Regulatory Potential.
Guanjue Xiang calculated the beta-coefficients and esRP scores. Belinda Giardine generated the track displayed and developed the track hub.
iGregory T, Yu C, Ma A, Orkin SH, Blobel GA, Weiss MJ. GATA-1 and erythropoietin cooperate to promote erythroid cell survival by regulating bcl-xL expression. Blood. 1999; 94:87-96. PMID: 10381501.
Pinto do O P, Richter K, Carlsson L. Hematopoietic progenitor/stem cells immortalized by Lhx2 generate functional hematopoietic cells in vivo. Blood. 2002 Jun 1;99(11):3939-46. doi: 10.1182/blood.v99.11.3939. PMID: 12010792.
Weiss MJ, Yu C, Orkin SH. Erythroid-cell-specific properties of transcription factor GATA-1 revealed by phenotypic rescue of a gene-targeted cell line. Mol Cell Biol. 1997; 17:1642-1651. PMID: 9032291; PMCID: PMC231889.
Xiang G, Keller CA, Heuston E, Giardine BM, An L, Wixom AQ, Miller A, Cockburn A, Sauria MEG, Weaver K, Lichtenberg J, Göttgens B, Li Q, Bodine D, Mahony S, Taylor J, Blobel GA, Weiss MJ, Cheng Y, Yue F, Hughes J, Higgs DR, Zhang Y, Hardison RC. An integrative view of the regulatory and transcriptional landscapes in mouse hematopoiesis. Genome Res. 2020 Mar;30(3):472-484. PMID: 32132109; PMCID: PMC7111515.
Xiang G, He X, Giardine BM, Isaac KJ, Taylor DJ, McCoy RC, Jansen C, Keller CA, Wixom AQ, Cockburn A, Miller A, Qi Q, He Y, Li Y, Lichtenberg J, Heuston EF, Anderson SM, Luan J, Vermunt MW, Yue F, Sauria MEG, Schatz MC, Taylor J, Göttgens B, Hughes JR, Higgs DR, Weiss MJ, Cheng Y, Blobel GA, Bodine DM, Zhang Y, Li Q, Mahony S, Hardison RC. Interspecies regulatory landscapes and elements revealed by novel joint systematic integration of human and mouse blood cell epigenomes. Genome Res. 2024 Aug 20;34(7):1089-1105. PMID: 38951027; PMCID: PMC11368181.
These data are available for use without restrictions.
Ross Hardison rch8@psu.edu